The Oppermann Lab
Welcome!

from left: Hong Yan, Paul Herkert, Laura Isigkeit, Prof. Sina Oppermann, Rekia Sinderwald, Catarina Santos Carvalho, Jasmina Neumann, Johanna Saumer
Forschung/ Research
Forschung/ Research
Funktionell-zielgerichtete Präzisionsmedizin bei Krebspatienten - von präklinischen ex vivo Studien zur klinischen Anwendung
Ziel der Präzisionsmedizin (PM) ist es, Behandlungsstrategien auf die Bedürfnisse eines einzelnen Patienten zuzuschneiden. Bisher basiert die personalisierte Krebsbehandlung weitgehend auf genomischen Tumorprofilen, die für viele Patienten unbefriedigende Ergebnisse liefern. Unsere Arbeitsgruppe „Funktionelle Präzisionsonkologie“ hat das Ziel, die genomischen Ansätze durch eine funktionelle ex-vivo Wirkstoffsensitivitäts- und Resistenztestung (engl: Drug Response Profiling, DRP) unter Verwendung von patienteneigenen Kurzzeit-Tumorkulturen zu ergänzen.
Dabei stehen drei Ziele im Vordergrund:
1) die Untersuchung von krebs- und wirkstoffinduzierten Phänotypen, um neue Wirkstoffe und Möglichkeiten zur Neupositionierung von Medikamenten zu entdecken;
2) die Untersuchung möglicher Biomarker für die beobachteten Medikamentenwirkungen sowie für die entstehenden Resistenzen und
3) die direkte Übertragung der Ergebnisse in die Klinik.
Unser stark kollaborativer Ansatz fördert ein starkes Netzwerk zwischen verschiedenen Disziplinen, insbesondere zwischen Pharmazie und Medizin sowie (klinischer) Bioinformatik, und stärkt dadurch die interprofessionelle Zusammenarbeit.
Functional Precision Oncology
The concept of precision oncology (PM) is being used worldwide to identify new therapy strategies for individual patients, mainly for high-risk patients who suffer from uncurable, refractory or relapsed aggressive diseases. State-of-the art next-generation sequencing techniques are used to identify the drug that efficiently works for the respective patient by targeting the tumor-driving genetic aberration(s) (e.g., fusion, mutation, or amplification) present in the individual tumor cells (Letai et al., 2017). However, it has become apparent that sequencing of the biopsy sample alone is not sufficient to assign a tailor-made drug for each patient and not all patients do currently benefit from therapy solely selected based on the molecular target identification (Kornauth et al., 2022).
To address these challenges of current genomic PM programs and to bridge the gap between the genomic knowledge of individual cancers and what can be directly applied in the clinic, comprehensive functional information is needed to identify the vulnerability of the respective tumor.
Our group functional precision oncology is addressing this need by performing functional ex vivo drug efficacy testing on patient-derived tumor cells to complement genomic approaches, particularly for cases in which genomics fail to guide treatments or in which the tumor is driven by non-genomic factors, such as the tumor microenvironment.
By providing functional information at the individual tumor cell level, we use aim to
1) better understand cancer- and drug-induced phenotypes to discover new drug efficacies and drug repositioning opportunities;
2) to investigate possible biomarkers and underlying mechanisms for the drug effects seen as well as for the resistances that emerge, and
3) to ultimately translate the results directly into the clinic.
To these aims we are carrying out the following projects:
- Predictive ex vivo cancer models
- Functional drug response testing
- High-content phenotypical image analysis using artificial intelligence
As the ultimate goal is to implement the functional precision medicine approach into a widespread clinical use, we aim for a high standardization of our functional assays to ensure reproducibility and reliability or our protocols and results. To this aim we are collaborating internationally with many of clinical and academic research groups in the field of FPM sharing protocols and knowledge. For clinical use our assays are further developed in a way that allows a quick turnaround time of results allowing a clinical decision making within a maximum of a couple of weeks. We aim to use our results for the design of new clinical trials further allowing for clinical validation. Last, we are addressing current challenges, such as regulatory and reimbursement issues as well as cost and accessibility, needed for a broader use of functional precision cancer medicine.
Methoden/ Methods
Methoden/ Methods
Die Schlüsselmethodik der Plattform für funktionelle Präzisionsmedizin ist der Einsatz der konfokalen Hochdurchsatz- Mikroskopie (HCM, Bioimaging), die eine schnelle Testung zugelassener und experimenteller Wirkstoffbibliotheken ermöglicht, um personalisierte Therapien für einzelne Patienten mit bislang unheilbaren Erkrankungen zu entwickeln. Die multi-parametrische (high-content) Analyse unter Einsatz von maschinellem Lernen liefert darüber hinaus ein tieferes Verständnis über das Verhalten einzelner Tumorzellen, sowie über die zugrundeliegenden Mechanismen von Therapieansprechen und -resistenz. Durch Berücksichtigung von klinischen sowie pharmakokinetischen und -dynamischen Daten der Medikamente in unseren Analysen unterstützen wir die Vorhersage der Wirksamkeit und Sicherheit in der direkten klinischen Anwendung. Die Integration der funktionellen Daten mit verfügbaren genomischen und proteomischen Daten der jeweiligen Tumorproben der Patienten dient der Untersuchung von Biomarkern und der Entwicklung neuer klinischer Studien. Durch den Einsatz von künstlicher Intelligenz und die Entwicklung von automatisierten Analysepipelines für die Integration und Auswertung der großen Datenmengen soll nicht nur eine schnelle Turnaround Zeit für eine direkte klinische Translation der ex vivo Ergebnisse erreicht werden, sondern auch neue Erkenntnisse über das Therapieansprechen der Patienten gewonnen werden. In enger Kooperation mit den Kliniken und behandelnden Ärzten sollen Ergebnisse im Tumorboard besprochen und eine weitere Validierung des klinischen Benefits ermöglicht werden und damit der translationale Erfolg der ex vivo Forschung zu verbessert werden. Ultimativ, hat unsere Gruppe das Ziel, die derzeitigen molekularen und immuntherapeutischen Präzisionsmedizin-Programme durch die Implementierung funktioneller Informationen über das Ansprechen individueller Tumorzellen auf Wirkstoffe maßgeblich voranzubringen.
Establishment of predictive and advanced ex vivo disease models:
We receive fresh and fresh-frozen patient-derived tumor material, which is used to establish predictive disease models. Current main cancer entities cover adult and pediatric leukemia (CLL, AML) as well as selected solid tumor entities. Here, we focus on establishing ex vivo tumor models that closely reflect the situation in the human body and at the same time are suitable for high-content drug testing in multi-well formats. We investigating different culture protocols that particularly mimic the tumor microenvironment and the influence of cellular matrices, including 2D (co-)cultures as well as 3D spheroid cultures with emphasize on short-term models that allow a relevant clinical turn-around time of our drug testing results.
Short-term and long-term tumor models are further used to discover underlying mechanisms for the drug efficacies and resistance seen. In reverse translation studies, co-culture systems including stromal cells, immune cells and/or cytokine signaling are established (Oppermann et al., Blood, 2016) to particularly investigate possible factors leading to in vivo drug resistance.
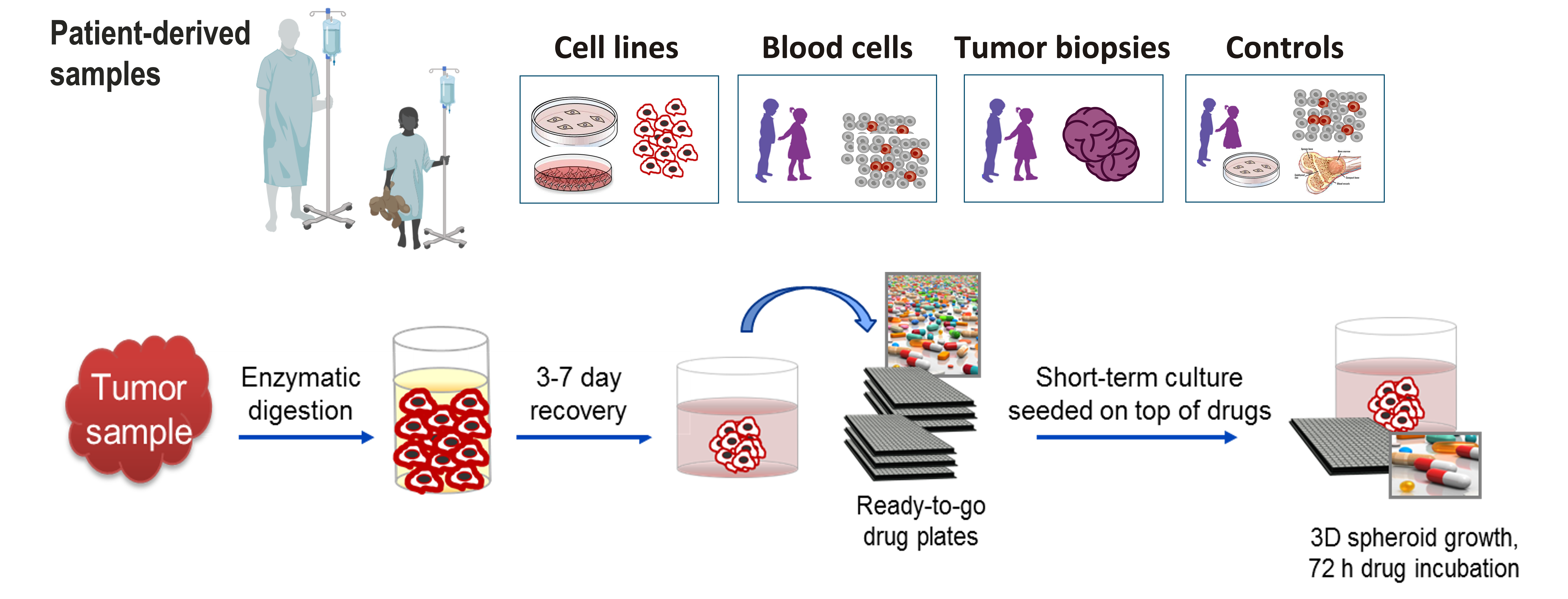
Figure: generated with biorender.com and further modified.
Functional drug response profiling:
We are carrying out functional ex vivo drug sensitivity and resistance profiling (‘drug response profiling’, DRP) using primary human material derived from cancer patients and healthy donors. Key methodology is the use of high-content fluorescent spinning- disc confocal microscopy (HCM, bioimaging) allowing to test large drug libraries for their effect on human-derived cancer cells in a medium- to high-throughput format.
We test different compound libraries, including donated chemical probe library obtained through the Structural Genomics Consortium (SGC) as well as customized drug libraries consisting mainly oncology drugs, that are FDA/EMA approved including conventional chemotherapeutic agents according to guideline and standard-of-care treatments for the respective diagnoses (see above), as well as a variety of targeted new drugs (e.g., (multi) kinase inhibitors) and compounds currently tested in clinical late phase (Phase II/III) for the respective diagnoses. For mechanistic studies and training controls, different cell death modulators (e.g., for apoptosis, necroptosis, ferroptosis and ER-stress-induced cell death, differential/epigenetic and metabolic modifiers) are included. Drugs are tested in a minimum of 5 concentrations covering the clinical achievable plasma and steady state concentration and replicates, mainly in a 384 multi-well format.
As automated analysis tools are crucial for handling and management of big data and to reach a quick turnaround time for clinical translation of results, we use and further develop bioinformatical drug analysis pipelines allowing the quantification of mono- and combination therapy responses for individual patients and disease cohorts and mapping of drug hits with drug target genes and multi-omics data (ElHarouni et al., Pharmacol Res., 2022).
Together with the molecular profiling of the respective tumor samples, the functional information will further used for establishment of hypotheses on cancer subtype-selective drug combinations and their predictive biomarkers. This synergistic approach ultimately also supports the optimal design of new clinical trials (e.g., new basket trials). Finally, functional measurements in combination with molecular cancer profiles provide valuable information for subsequent mechanistic studies aimed at validating potentially novel/drug effective targets
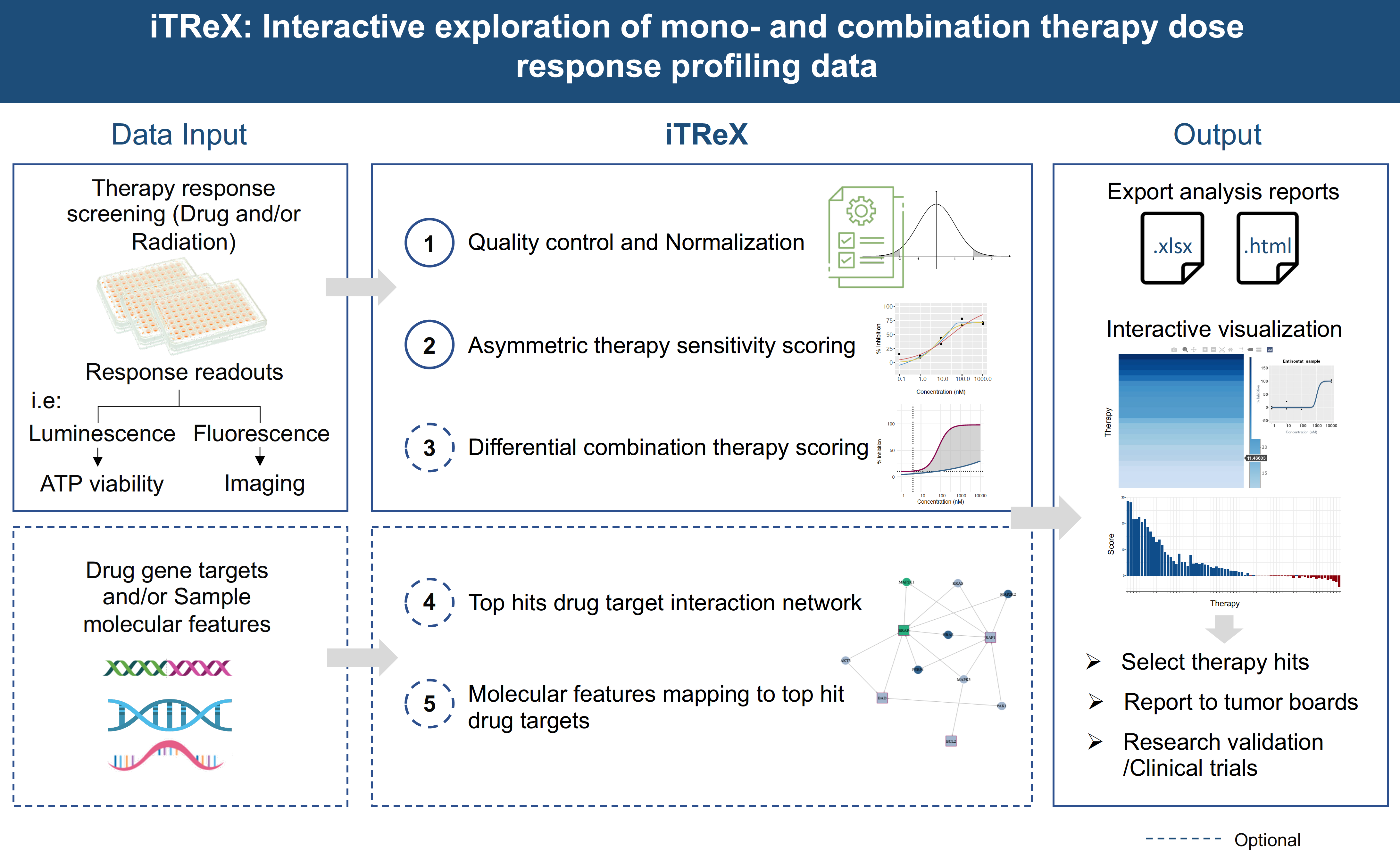
iTReX: ElHarouni et al., Pharmacological Research, 2022; https://itrex.kitz-heidelberg.de/
High-content phenotypical image analysis using artificial intelligence:
We apply high-content 2D and 3D image analysis to phenotypical drug response profiling of the patient-derived tumor cell cultures. Here we use the multiplex/cell painting approach, combining different fluorescent dyes to extract phenotypical information of single cells and applied drugs. Using this phenotypical profiling approach, we investigate how the different drugs affect the different subpopulations of tumor and non-tumor cells and further classify induced cell death mechanisms as well as entity and patient specific phenotypes and drug response. We employ both classical image analysis approaches (MIP (2D), volumetric 3D analysis, 3D and 2D nuclei and single cell segmentation, advanced feature extraction (i.e morphology, shape, intensity, texture), supervised and unsupervised phenotype clustering) as well as end-to-end deep learning with the aim to gain more complex and detailed information of drug response and cellular behavior.
For quantification of image-based drug responses, we have previously developed a patient-by-patient deep transfer learning method using a convolutional neural network which transfers image information into image-based cell viabilities (Berker Y. et al., IEEE-Trans Med Imaging (TMI), 2022). Our group is now building up on the established pipelines and is developing new and advanced AI-based image analysis models. Here, we closely collaborate with the local and international bioinformatics, AI and image-analysis groups.
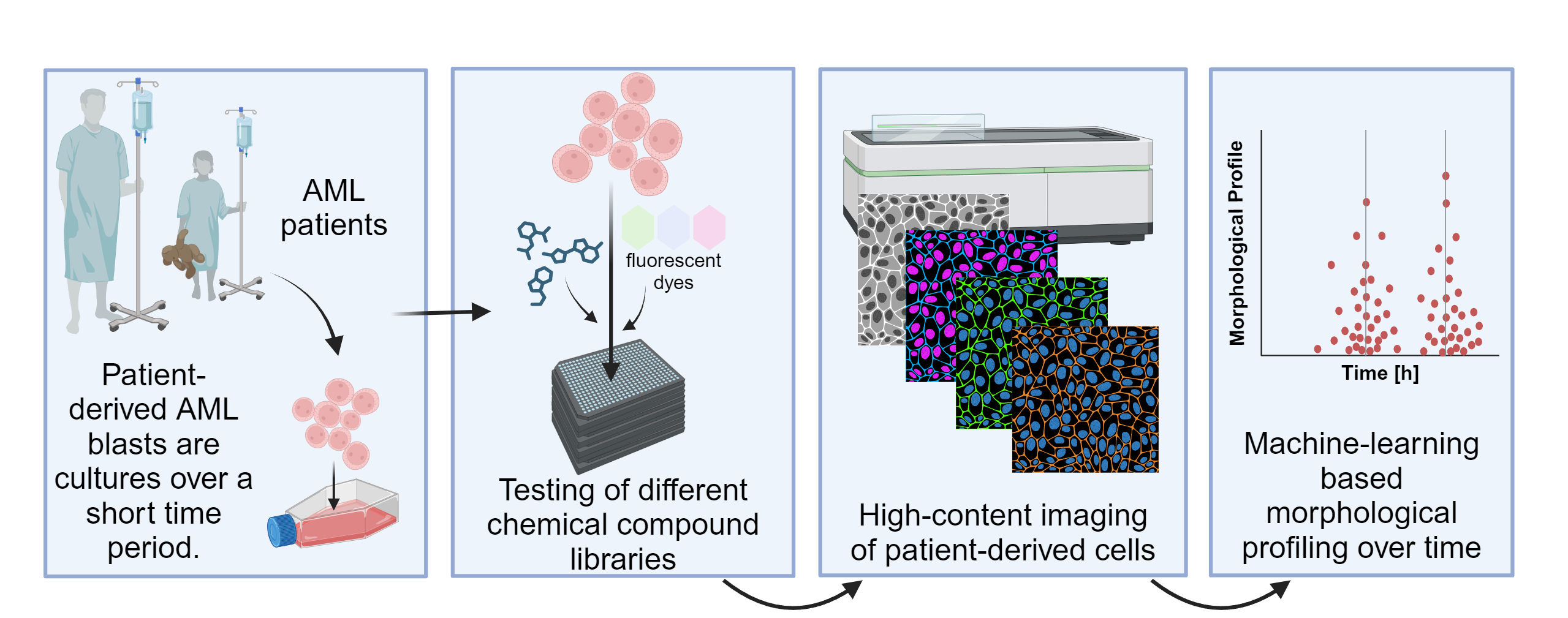
Figure: Example workflow for functional drug response testing on patient-derived AML blasts.
Publications
Publications
- Zeuner S, Vollmer J, Sigaud R, Oppermann S, Peterziel H, ElHarouni D, Oehme I, Witt O, Milde T, Ecker J. Combination drug screen identifies synergistic drug interaction of BCL-XL and class I histone deacetylase inhibitors in MYC-amplified medulloblastoma cells. J Neurooncol. 2024 Jan;166(1):99-112. doi: 10.1007/s11060-023-04526-w. Epub 2024 Jan 7. PMID: 38184819; PMCID: PMC10824805.
- Peterziel H, Jamaladdin N, ElHarouni D, Gerloff XF, Herter S, Fiesel P, Berker Y, Blattner-Johnson M, Schramm K, Jones BC, Reuss D, Turunen L, Friedenauer A, Holland-Letz T, Sill M, Weiser L, Previti C, Balasubramanian G, Gerber NU, Gojo J, Hutter C, Øra I, Lohi O, Kattamis A, de Wilde B, Westermann F, Tippelt S, Graf N, Nathrath M, Sparber-Sauer M, Sehested A, Kramm CM, Dirksen U, Kallioniemi O, Pfister SM, van Tilburg CM, Jones DTW, Saarela J, Pietiäinen V, Jäger N, Schlesner M, Kopp-Schneider A, Oppermann S, Milde T, Witt O, Oehme I. Drug sensitivity profiling of 3D tumor tissue cultures in the pediatric precision oncology program INFORM. NPJ Precis Oncol. 2022 Dec 27;6(1):94. doi: 10.1038/s41698-022-00335-y. PMID: 36575299; PMCID: PMC9794727.
- Berker Y, ElHarouni D, Peterziel H, Fiesel P, Witt O, Oehme I, Schlesner M, Oppermann S. Patient-by-Patient Deep Transfer Learning for Drug-Response Profiling Using Confocal Fluorescence Microscopy of Pediatric Patient-Derived Tumor-Cell Spheroids. IEEE Trans Med Imaging. 2022 Dec;41(12):3981-3999. doi: 10.1109/TMI.2022.3205554. Epub 2022 Dec 2. PMID: 36099221.
- Gatzweiler C, Ridinger J, Herter S, Gerloff XF, ElHarouni D, Berker Y, Imle R, Schmitt L, Kreth S, Stainczyk S, Ayhan S, Najafi S, Krunic D, Frese K, Meder B, Reuss D, Fiesel P, Schramm K, Blattner-Johnson M, Jones DTW, Banito A, Westermann F, Oppermann S, Milde T, Peterziel H, Witt O, Oehme I. Functional Therapeutic Target Validation Using Pediatric Zebrafish Xenograft Models. Cancers (Basel). 2022 Feb 8;14(3):849. doi: 10.3390/cancers14030849. PMID: 35159116; PMCID: PMC8834194.
- ElHarouni D, Berker Y, Peterziel H, Gopisetty A, Turunen L, Kreth S, Stainczyk SA, Oehme I, Pietiäinen V, Jäger N, Witt O, Schlesner M, Oppermann S. iTReX: Interactive exploration of mono- and combination therapy dose response profiling data. Pharmacol Res. 2022 Jan;175:105996. doi: 10.1016/j.phrs.2021.105996. Epub 2021 Nov 27. PMID: 34848323.
- Dietrich S, Oleś M, Lu J, Sellner L, Anders S, Velten B, Wu B, Hüllein J, da Silva Liberio M, Walther T, Wagner L, Rabe S, Ghidelli-Disse S, Bantscheff M, Oleś AK, Słabicki M, Mock A, Oakes CC, Wang S, Oppermann S, Lukas M, Kim V, Sill M, Benner A, Jauch A, Sutton LA, Young E, Rosenquist R, Liu X, Jethwa A, Lee KS, Lewis J, Putzker K, Lutz C, Rossi D, Mokhir A, Oellerich T, Zirlik K, Herling M, Nguyen-Khac F, Plass C, Andersson E, Mustjoki S, von Kalle C, Ho AD, Hensel M, Dürig J, Ringshausen I, Zapatka M, Huber W, Zenz T. Drug-perturbation-based stratification of blood cancer. J Clin Invest. 2018 Jan 2;128(1):427-445. doi: 10.1172/JCI93801. Epub 2017 Dec 11. PMID: 29227286; PMCID: PMC5749541.
- Oppermann S, Lam AJ, Tung S, Shi Y, McCaw L, Wang G, Ylanko J, Leber B, Andrews D, Spaner DE. Janus and PI3-kinases mediate glucocorticoid resistance in activated chronic leukemia cells. Oncotarget. 2016 Nov 8;7(45):72608-72621. doi: 10.18632/oncotarget.11618. PMID: 27579615; PMCID: PMC5341931.
- Oppermann S, Ylanko J, Shi Y, Hariharan S, Oakes CC, Brauer PM, Zúñiga-Pflücker JC, Leber B, Spaner DE, Andrews DW. High-content screening identifies kinase inhibitors that overcome venetoclax resistance in activated CLL cells. Blood. 2016 Aug 18;128(7):934-47. doi: 10.1182/blood-2015-12-687814. Epub 2016 Jun 13. PMID: 27297795; PMCID: PMC5000578.
- Neitemeier S, Jelinek A, Laino V, Hoffmann L, Eisenbach I, Eying R, Ganjam GK, Dolga AM, Oppermann S, Culmsee C. BID links ferroptosis to mitochondrial cell death pathways. Redox Biol. 2017 Aug;12:558-570. doi: 10.1016/j.redox.2017.03.007. Epub 2017 Mar 9. PMID: 28384611; PMCID: PMC5382034.
- Brahmbhatt H, Oppermann S, Osterlund EJ, Leber B, Andrews DW. Molecular Pathways: Leveraging the BCL-2 Interactome to Kill Cancer Cells—Mitochondrial Outer Membrane Permeabilization and Beyond. Clin Cancer Res. 2015 Jun 15;21(12):2671-6. doi: 10.1158/1078-0432.CCR-14-0959. Epub 2015 Apr 2. PMID: 25838396.
Aktuelles/ News
Aktuelles/ News
Ein Trainingsraum für angehende Apotheker*innen
Bei Übungen in der Trainingsapotheke und mit dem Onlinetool »MyDispense« können Pharmazie-Studierende ihre Beratungs- und Fachkompetenz testen und erweitern.

Erschienen in: https://www.unireport.info/176087817/unireport-ausgabe-4-2025-vom-17-juli-2025.pdf (Seite 8)
Jahresbericht 2025 des Georg-Speyer-Hauses
Link: Functional Precision Oncology (Oppermann Lab)
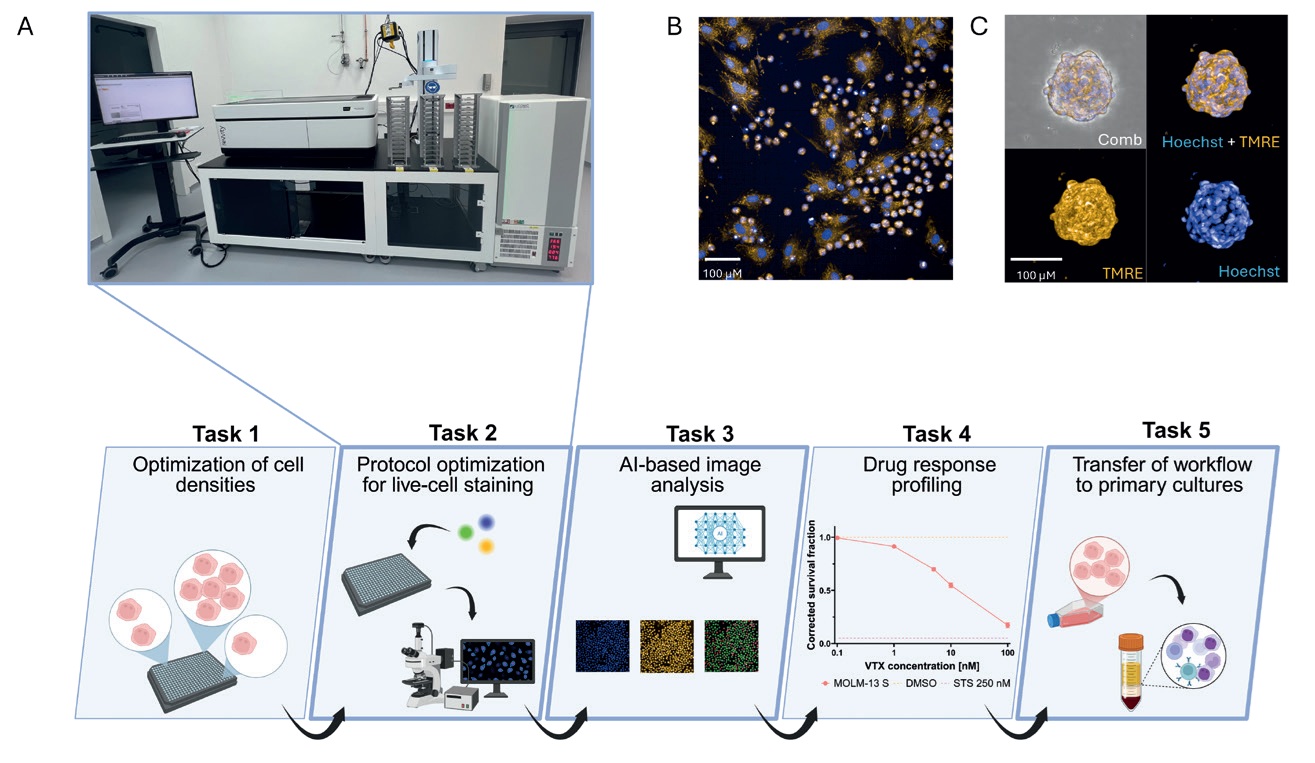
Figure: Optimization and translation of an image-based workflow enabling drug response profiling in AML. (A) Confocal spinning-disc microscopy (Opera Phenix Plus, Revvity). (B) Co-culture of MSCs and AML cells imaged by Opera Phenix; nuclei and mitochondrial potential were visualized using Hoechst (blue) and TMRE (orange), respectively. (C) Representative MSC spheroids imaged by Opera Phenix, stained with Hoechst (blue) and TMRE (orange).
Jobs
Jobs
Die Arbeitsgruppe von Frau Prof. Dr. Sina Oppermann am Institut für Pharmakologie und Klinische Pharmazie (Fachbereich Biochemie, Chemie und Pharmazie) sucht ab sofort:
zwei Studentische Hilfskräfte / Werkstudenten mit abgeschlossenem 2. Staatsexamen (Pharmazie)
(20-40 Std./Monat)
zur Unterstützung in der Lehre und Forschung.
Folgende Aufgaben kommen auf Sie zu:
- Unterstützung in der Lehre der Fächer Klinische Pharmazie und Krankheitslehre
- Erstellung und Bewertung von Patientenfällen für die Präsenz-Trainingsapotheke
- Mitwirkung beim Aufbau und der Pflege der virtuellen Apotheke „MyDispense“
- Organisation und Koordination des Wahlpflichtpraktikums Klinische Pharmazie
- Unterstützung in der Zellkultur und Durchführung einfacher Laborarbeiten
Voraussetzungen:
- Pharmazeut*in mit abgeschlossenem 2. Staatsexamen
- optimal im Master „Arzneimittelforschung“ (HiWi) oder im Praktischen Jahr in einer anderen Arbeitsgruppe des Fachbereichs (Werkstudent)
- Idealerweise vor Ort an der Goethe-Universität oder umliegenden Orten zur flexiblen Einsetzbarkeit vor Ort sowie Remote Verfügbarkeit
- Gute Kenntnisse in der Pharmakologie und Klinischen Pharmazie
- Sehr gute PC-Kenntnisse (inkl. MS Office und Grafikprogramme)
- Strukturierte, detaillierte und verlässliche Arbeitsweise
Ihre Bewerbung (Lebenslauf, Bewerbungsschreiben, aktuelle Studienbescheinigung) senden Sie bitte per E-Mail an Prof. Dr. Sina Oppermann, Institut für Pharmakologie und Klinische Pharmazie, Goethe-Universität, Max-von-Laue-Str. 9, 60438 Frankfurt am Main